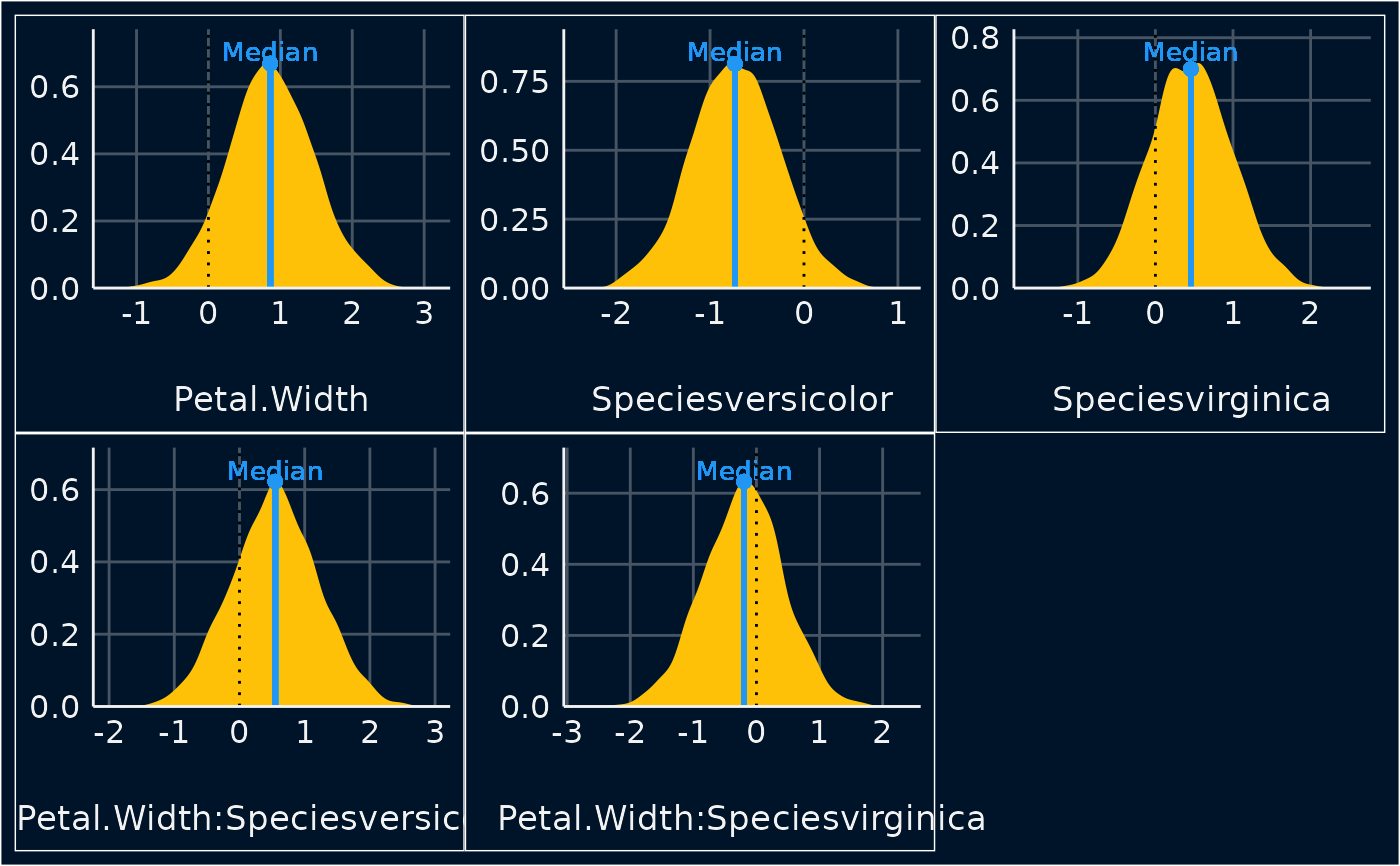
Plot method for point estimates of posterior samples
Source:R/plot.point_estimates.R
plot.see_point_estimate.RdThe plot() method for the bayestestR::point_estimate().
Usage
# S3 method for class 'see_point_estimate'
plot(
x,
data = NULL,
size_point = 2,
size_text = 3.5,
panel = TRUE,
show_labels = TRUE,
show_intercept = FALSE,
priors = FALSE,
alpha_priors = 0.4,
...
)Arguments
- x
An object.
- data
The original data used to create this object. Can be a statistical model.
- size_point
Numeric specifying size of point-geoms.
- size_text
Numeric value specifying size of text labels.
- panel
Logical, if
TRUE, plots are arranged as panels; else, single plots are returned.- show_labels
Logical. If
TRUE, the text labels for the point estimates (i.e. "Mean", "Median" and/or "MAP") are shown. You may setshow_labels = FALSEin case of overlapping labels, and add your own legend or footnote to the plot.- show_intercept
Logical, if
TRUE, the intercept-parameter is included in the plot. By default, it is hidden because in many cases the intercept-parameter has a posterior distribution on a very different location, so density curves of posterior distributions for other parameters are hardly visible.- priors
Logical. If
TRUE, prior distributions are simulated (usingbayestestR::simulate_prior()) and added to the plot.- alpha_priors
Numeric value specifying alpha for the prior distributions.
- ...
Arguments passed to or from other methods.
Examples
library(rstanarm)
library(bayestestR)
set.seed(123)
m <<- suppressWarnings(stan_glm(Sepal.Length ~ Petal.Width * Species, data = iris, refresh = 0))
result <- point_estimate(m, centrality = "median")
result
#> Point Estimate
#>
#> Parameter | Median
#> --------------------------------------
#> (Intercept) | 4.79
#> Petal.Width | 0.86
#> Speciesversicolor | -0.74
#> Speciesvirginica | 0.46
#> Petal.Width:Speciesversicolor | 0.55
#> Petal.Width:Speciesvirginica | -0.20
plot(result)