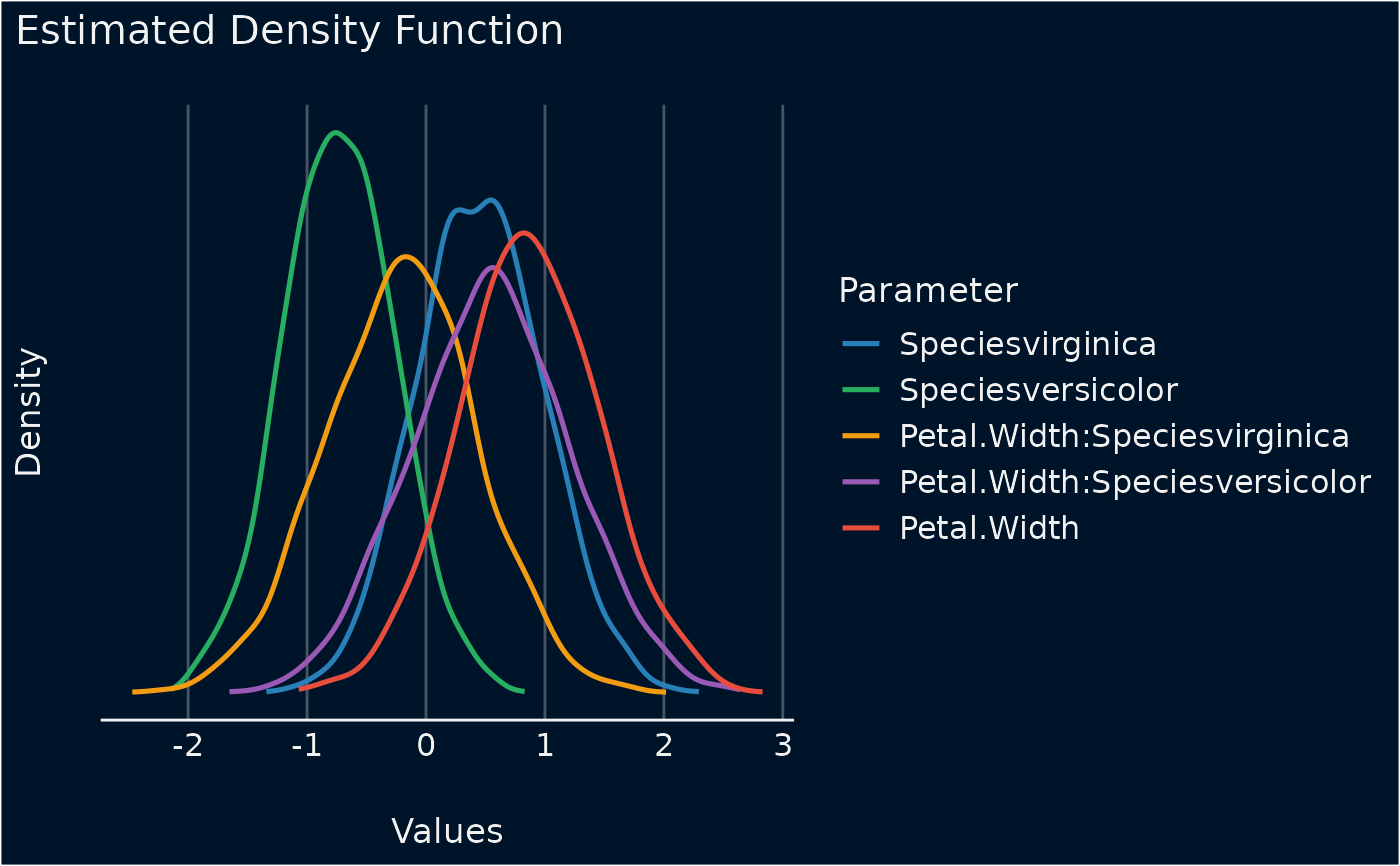
Plot method for density estimation of posterior samples
Source:R/plot.estimate_density.R
plot.see_estimate_density.RdThe plot() method for the bayestestR::estimate_density() function.
Usage
# S3 method for class 'see_estimate_density'
plot(
x,
stack = TRUE,
show_intercept = FALSE,
n_columns = 1,
priors = FALSE,
alpha_priors = 0.4,
alpha_posteriors = 0.7,
linewidth = 0.9,
size_point = 2,
centrality = "median",
ci = 0.95,
...
)Arguments
- x
An object.
- stack
Logical. If
TRUE, densities are plotted as stacked lines. Else, densities are plotted for each parameter among each other.- show_intercept
Logical, if
TRUE, the intercept-parameter is included in the plot. By default, it is hidden because in many cases the intercept-parameter has a posterior distribution on a very different location, so density curves of posterior distributions for other parameters are hardly visible.- n_columns
For models with multiple components (like fixed and random, count and zero-inflated), defines the number of columns for the panel-layout. If
NULL, a single, integrated plot is shown.- priors
Logical. If
TRUE, prior distributions are simulated (usingbayestestR::simulate_prior()) and added to the plot.- alpha_priors
Numeric value specifying alpha for the prior distributions.
- alpha_posteriors
Numeric value specifying alpha for the posterior distributions.
- linewidth
Numeric value specifying size of line geoms.
- size_point
Numeric specifying size of point-geoms.
- centrality
Character specifying the point-estimate (centrality index) to compute. Can be
"median","mean"or"MAP".- ci
Numeric value of probability of the CI (between 0 and 1) to be estimated. Default to
0.95.- ...
Arguments passed to or from other methods.
Examples
library(rstanarm)
library(bayestestR)
set.seed(123)
m <<- suppressWarnings(stan_glm(Sepal.Length ~ Petal.Width * Species, data = iris, refresh = 0))
result <- estimate_density(m)
plot(result)
#> Ignoring unknown labels:
#> • fill : "Parameter"