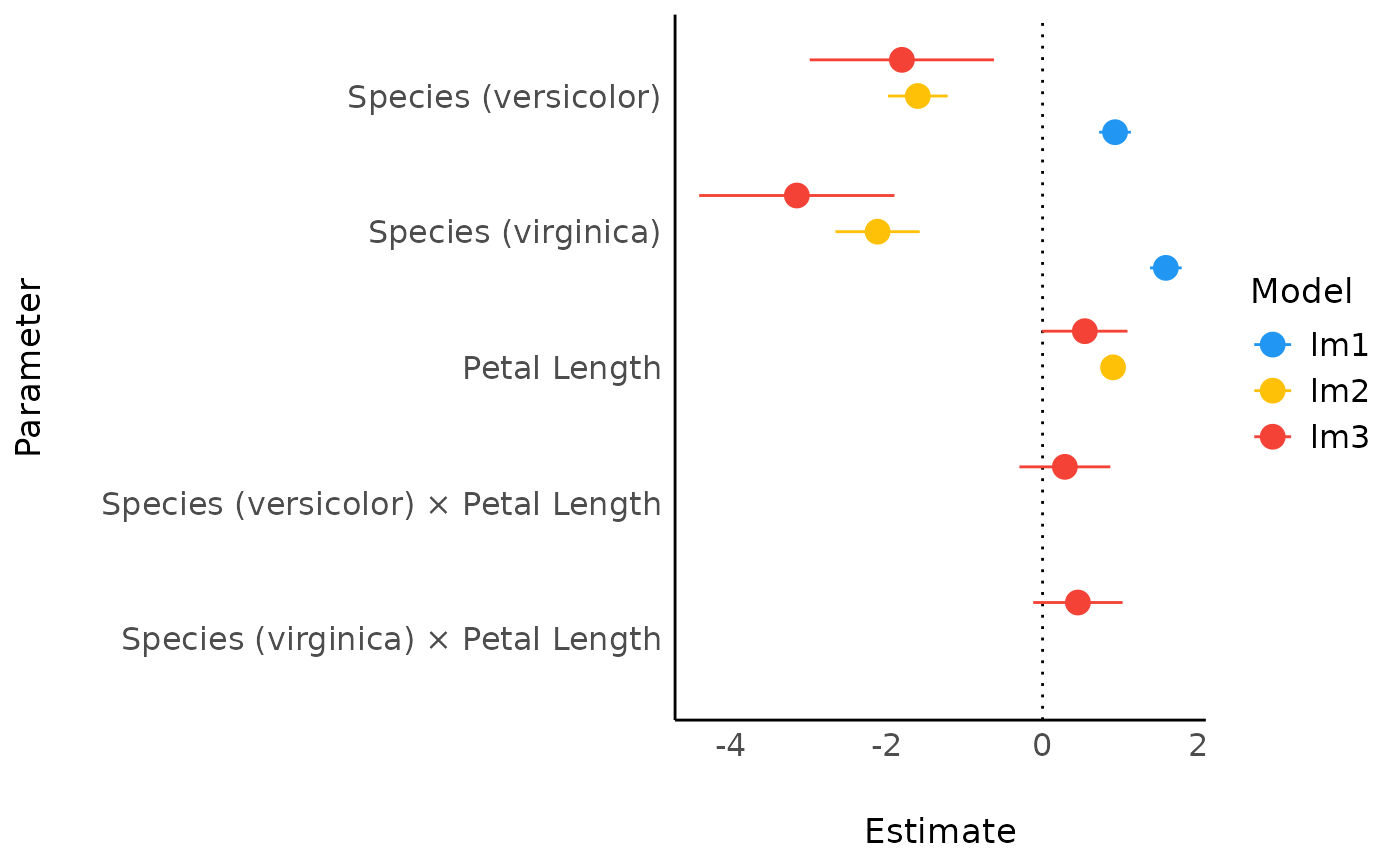
Plot method for comparison of model parameters
Source:R/plot.compare_parameters.R
plot.see_compare_parameters.RdThe plot() method for the parameters::compare_parameters()
function.
Usage
# S3 method for class 'see_compare_parameters'
plot(
x,
show_intercept = FALSE,
size_point = 0.8,
size_text = NA,
dodge_position = 0.8,
sort = NULL,
n_columns = NULL,
show_labels = FALSE,
...
)Arguments
- x
An object.
- show_intercept
Logical, if
TRUE, the intercept-parameter is included in the plot. By default, it is hidden because in many cases the intercept-parameter has a posterior distribution on a very different location, so density curves of posterior distributions for other parameters are hardly visible.- size_point
Numeric specifying size of point-geoms.
- size_text
Numeric value specifying size of text labels.
- dodge_position
Numeric value specifying the amount of "dodging" (spacing) between geoms.
- sort
The behavior of this argument depends on the plotting contexts.
Plotting model parameters: If
NULL, coefficients are plotted in the order as they appear in the summary. Settingsort = "ascending"orsort = "descending"sorts coefficients in ascending or descending order, respectively. Settingsort = TRUEis the same assort = "ascending".Plotting Bayes factors: Sort pie-slices by posterior probability (descending)?
- n_columns
For models with multiple components (like fixed and random, count and zero-inflated), defines the number of columns for the panel-layout. If
NULL, a single, integrated plot is shown.- show_labels
Logical. If
TRUE, text labels are displayed.- ...
Arguments passed to or from other methods.